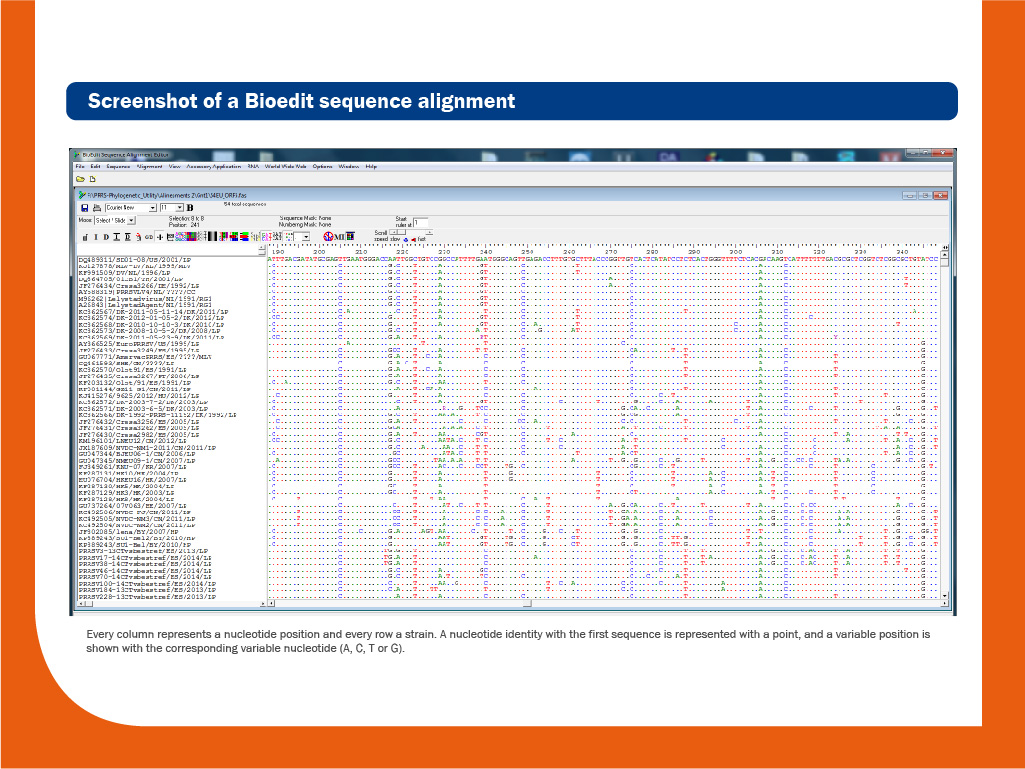
DNA obtained during the RT-PCR assay can be sequenced to obtain the order of nucleotides in the genome. DNA sequencing is useful to discriminate between individual strains. Usually, the relatedness among different isolates is done based on the comparison of ORF5, ORF6 or ORF7 sequences.
Regarding individual blood samples, it is important to note that Ct values below 32 are needed for sequencing; samples with Ct values > 32 are difficult to sequence since the amount of the virus is insufficient.
Oral fluids and pools of sera should be avoided for sequencing because they can easily contain more than one strain and because by mixing different samples we are diminishing the amount of virus present.
Sequencing is a complementary diagnostic tool. It is helpful to take decisions about measures to control the virus.
Through sequencing it is possible to:
- Distinguish species
- Distinguish field-virus isolates from vaccines
- Obtain valuable epidemiologic information useful to determine the source of infection, how many strains are present in a herd/farm/population, which is the most prevalent, recombination events, etc.
In general, if sequences are frequently monitored (it is recommended from each strain involved in each outbreak or periodically, at least every six months), we can determine the main source of infections and how isolates circulate in one herd or in a region.
Eventually, we could use this information to establish better measures for PRRS control.
Through sequencing it is not possible to:
- Predict the virulence of a given isolate, in general.
- Predict the level of cross-protection between two isolates, since genetic markers of virulence and cross-protection have not been properly identified.
Sequencing should not be used to decide which vaccine should be used against a given isolate, since it has been demonstrated that the level of cross-protection that could exist between two strains is not related to the overall level of homology (relatedness) between them.
In order to compare the degree of similarity among isolates and to group them according to their sequences, a phylogenetic tree can be drawn.
Trees are more useful to illustrate relationships among various isolates than just a comparison of percentages of homology.
It may happen that a group of strains can share the same percentage of similitude even being different among them; in these cases, trees are crucial to observe the real relationships.
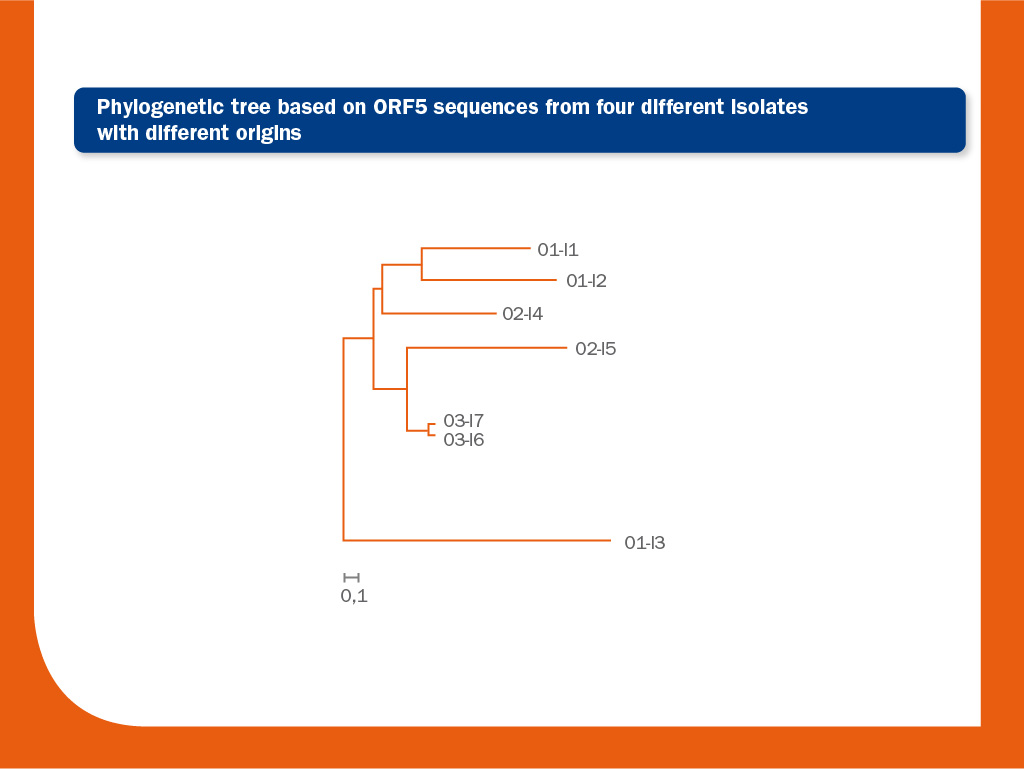
The image above represents a phylogenetic tree obtained by the analysis of seven ORF5 sequences from four different origins.
The bar represents 0.1 nucleotide substitution per site. The interpretation of this tree is as follows: in the lower branch one unrelated strain from origin 1 (O1) is present (O1-I3) (similarity to O1-I1 and O1-I2 from the same origin, located in the upper branch of the tree is 93.4% and 93.2%, respectively).
Similarity between these two related strains (O1-I1 and O1-I2) is 97.5%. In origin 2, two unrelated strains I4 and I5 are also present (similarity between them is 96.6%).
Probably, there is a link between O2-I4 and the origin 1 and between O2-I5 and origin 3. Finally, the strains I6 and I7 from origin 3 are closely related (>99.8% identical).
It is commonly accepted that two isolates are different if similarity between them is below 97% (ORF5).
However, this cut-off point is completely arbitrary. In order to avoid misinterpretations, additional information, such as dates and locations of isolations, as well as the relationships among farms or origins, should be taken into account.
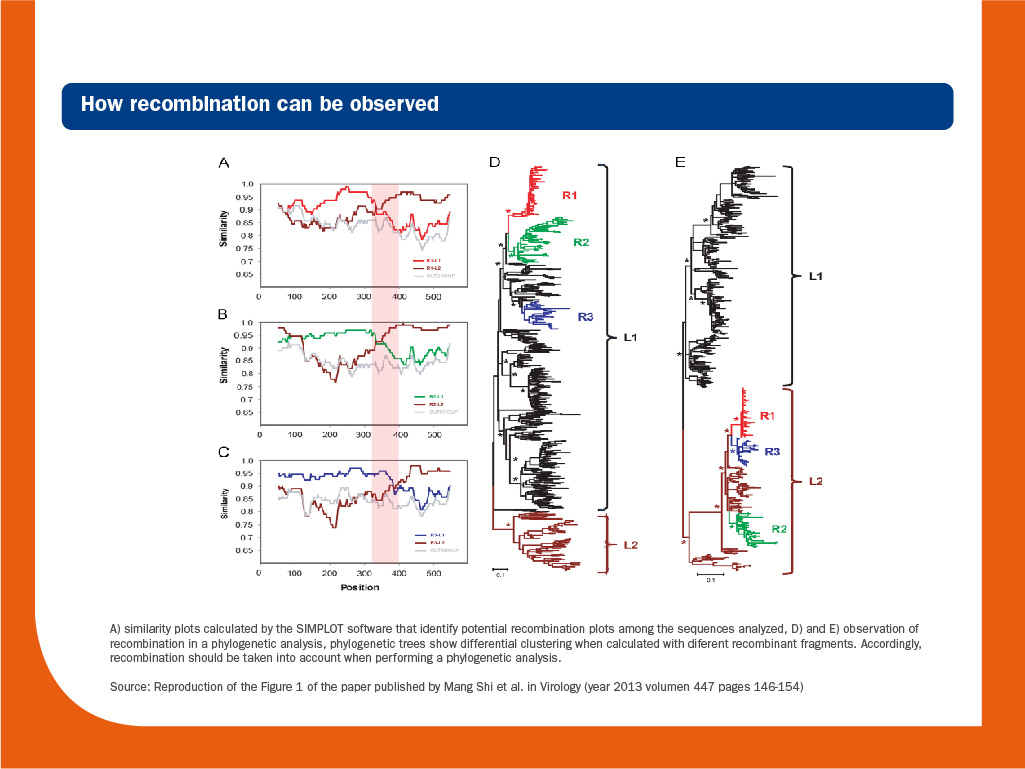
It is worth to note that recombination should be considered when phylogenetic trees are calculated, since it is a hallmark of RNA virus evolution and can blur the true phylogenetic relationships among strains.
Sanger sequencing
Until recently, Sanger sequencing has been the main tool to study genetic diversity. However, though useful this technique has several limitations; it only retrieves consensus sequences, the most common variant present in a sample.
As it has been repeatedly stated, PRRS virus exists in the host as quasispecies. Therefore, low frequency mutants are neglected when Sanger is used.
Moreover, when two or more different strains are present, sequences obtained by Sanger technique are very difficult to interpret.
Although it represents a very limited proportion of the total genome (4%), ORF5 is usually used to compare strains, since it has a high mutation rate and changes are easier to observe.
Obviously, whole genome sequencing would yield much more information:
- More accurately phylogenetic grouping and comparisons among strains.
- Identification of potential epitopes, epidemiological or virulence markers.
- Identification of recombination events beyond ORF5 or ORF7.
However, whole genome Sanger sequencing it is a very time-consuming task (it requires the sequencing of many consecutive overlapping genomes segments amplified previously by PCR).
Consequently, compared to ORF5 or ORF7, few whole genome sequences are available.
Next-generation sequencing
Several of the limitations of Sanger sequencing can be overcome using Next-generation sequencing (NGS), also known as high-throughput sequencing. The term is used to describe a number of different modern non-Sanger-based sequencing technologies.
These technologies are very powerful and certainly a revolution in the field of genomics and molecular biology, since they allow us to sequence DNA and RNA much more quickly and extremely deeper than Sanger sequencing.
Advantages of NGS:
- Easiness to obtain full-genome sequences.
- Easiness to determine if more than one strain is present in sample.
- Easiness to identify recombination events.
- Full characterization of the mutation swarm. It is possible to obtain from hundred to hundreds of thousands of sequences from just one sample.
Limitations of NGS:
- The huge data obtained from just one sample can be difficult to manage.
- High skills in bioinformatics are required to evaluate sequence reads and to minimize misinterpretation of data.
- Difficulty discerning if a sequence read showing to a low-frequency mutation has a biological significance.
- Until now, the cost.
© Laboratorios Hipra, S.A. 2024. All Rights Reserved.
No part of this website or any of its contents may be reproduced, copied, modified or adapted, without the prior written consent of HIPRA.
- Bautista EM, Meulenberg JJ, Choi CS, Molitor TW. Structural polypeptides of the American (VR-2332) strain of porcine reproductive and respiratory syndrome virus. Arch Virol. 1996:1357-65.
- Benfield DA, Nelson E, Collins JE, Harris L, Hennings JC, Shaw DP, Goyal SM, McCullough S, Morrison RB, Joo HS, Gorcyca D, Chladek D. Characterization of swine infertility and respiratory syndrome (SIRS) virus (isolate ATCC VR-2332). J Vet Diagn Invest. 1992. 4, 127-133.
- Benfield D, Nelson J, Rossow K, Nelson C, Steffen M, Rowland R. Diagnosis of persistent or prolonged porcine reproductive and respiratory syndrome virus infections Vet Res. 2000, 31:71.
- Brockmeier SL, Halbur PG, Thacker EL. Porcine respiratory disease complex. In KA Brogden, JM Guthmiller, eds, Polymicrobial Diseases. Washington, DC: ASM Press. 2002, 231-58.
- Calzada-Nova G, Schnitzlein W, Husmann R, Zuckermann FA. Characterization of the cytokine and maturation responses of pure populations of porcine plasmacytoid dendritic cells to porcine viruses and toll-like receptor agonists. Vet Immunol Immunopathol. 2010, 135:20-33.
- Chen WY, Schniztlein WM, Calzada-Nova G, Zuckermann FA. Genotype 2 strains of porcine reproductive and respiratory syndrome virus dysregulate alveolar macrophage cytokine production via the unfolded protein response. J Virol.2017, pii: JVI.01251-17.
- Christopher-Hennings J, Nelson EA, Hines RJ, Nelson JK, Swenson SL, Zimmerman JJ, Chase CL, Yaeger MJ, Benfield DA. Persistence of porcine reproductive and respiratory syndrome virus in serum and semen of adult boars. J Vet Diagn Invest. 1995, 7:456-64.
- Cortey M, Díaz I, Martín-Valls GE, Mateu E. Next-generation sequencing as a tool for the study of Porcine reproductive and respiratory syndrome virus (PRRSV) macro- and micro- molecular epidemiology. Vet Microbiol. 2017. doi: 10.1016/j.vetmic.2017.02.002.
- Darwich L, Díaz I, Mateu E. Certainties, doubts and hypotheses in porcine reproductive and respiratory syndrome virus immunobiology. Virus Res. 2010, 154:123-32.
- Díaz I, Darwich L, Pappaterra G, Pujols J, Mateu E. Different European-type vaccines against porcine reproductive and respiratory syndrome virus have different immunological properties and confer different protection to pigs. Virology. 2006, 351:249-59.
- Díaz I, Gimeno M, Darwich L, Navarro N, Kuzemtseva L, López S, Galindo I, Segalés J, Martín M, Pujols J, Mateu E. Characterization of homologous and heterologous adaptive immune responses in porcine reproductive and respiratory syndrome virus infection. Vet Res. 2012, 19:43:30.
- García-Nicolás O, Quereda JJ, Gómez-Laguna J, Salguero FJ, Carrasco L, Ramis G, Pallarés FJ. Cytokines transcript levels in lung and lymphoid organs during genotype 1 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) infection. Vet Immunol Immunopathol. 2014, 160:26-40.
- Gimeno M, Darwich L, Diaz I, de la Torre E, Pujols J, Martín M, Inumaru S, Cano E, Domingo M, Montoya M, Mateu E. Cytokine profiles and phenotype regulation of antigen presenting cells by genotype-I porcine reproductive and respiratory syndrome virus isolates. Vet Res. 2011, 42:9.
- Gómez-Laguna J, Salguero FJ, Pallarés FJ, Carrasco L. Immunopathogenesis of porcine reproductive and respiratory syndrome in the respiratory tract of pigs. Vet J. 2013, 195:148-55.
- Haynes JS, Halbur PG, Sirinarumitr T, Paul PS, Meng XJ, Huffman EL. Temporal and morphologic characterization of the distribution of porcine reproductive and respiratory syndrome virus (PRRSV) by in situ hybridization in pigs infected with isolates of PRRSV that differ in virulence. Vet Pathol. 1997, 34:39-43.
- Hill H. Overview and history of Mystery Swine Disease (swine infertility/respiratory syndrome). Proceedings of the Mystery Swine Disease Committee Meeting, Livestock Conversation Institute, Denver, CO. 1990, 29–31.
- Holtkamp DJ, Polson DD, Torremorrell M. Terminology for classifying swine herds by porcine reproductive and respiratory status. J Swine Health and Prod. 2011, 19:44-56.
- Horter DC, Pogranichniy RM, Chang CC, Evans RB, Yoon KJ, Zimmerman JJ. Characterization of the carrier state in porcine reproductive and respiratory syndrome virus infection. Vet Microbiol. 2002, 86:213-28.
- Loula T. Mystery pig disease. Agri-practice. 1991, 12:23–34.
- Lunney JK, Benfield DA, Rowland RR. Porcine reproductive and respiratory syndrome virus: an update on an emerging and re-emerging viral disease of swine. Virus Res. 2010, 154:1-6.
- Mateu E, Tello M, Coll A, Casal J, Martín M. Comparison of three ELISAs for the diagnosis of porcine reproductive and respiratory syndrome. Vet Rec. 2006, 159:717-8.
- Mateu E, Diaz I. The challenge of PRRS immunology. Vet J. 2008, 177:345-51.
- Meier WA, Galeota J, Osorio FA, Husmann RJ, Schnitzlein WM, Zuckermann FA. Gradual development of the interferon-gamma response of swine to porcine reproductive and respiratory syndrome virus infection or vaccination. Virology. 2003, 309:18-31.
- Nelson EA, Christopher-Hennings J, Benfield DA. Serum immune responses to the proteins of porcine reproductive and respiratory syndrome (PRRS) virus. J Vet Diagn Invest. 1994, 6:410-5.
- Appendices IV and V of the REPORT OF THE OIE AD HOC GROUP ON PORCINE REPRODUCTIVE RESPIRATORY SYNDROME Paris, 9 – 11 June 2008 http://www.oie.int/fileadmin/Home/eng/Our_scientific_expertise/docs/pdf/PRRS_guide_web_bulletin.pdf.
- Rodríguez-Gómez IM, Gómez-Laguna J, Carrasco L. Impact of PRRSV on activation and viability of antigen presenting cells. World J Virol. 2013, 2:146-51.
- Rovira A, Cano JP, Muñoz-Zanzi C. Feasibility of pooled-sample testing for the detection of porcine reproductive and respiratory syndrome virus antibodies on serum samples by ELISA. Vet Microbiol. 2008, 130:60-8.
- Rovira A, Clement T, Christopher-Hennings J, Thompson B, Engle M, Reicks D, Muñoz-Zanzi C. Evaluation of the sensitivity of reverse-transcription polymerase chain reaction to detect porcine reproductive and respiratory syndrome virus on individual and pooled samples from boars. J Vet Diagn Invest. 2007, 19:502-9.
- Rovira A, Reicks D, Muñoz-Zanzi C. Evaluation of surveillance protocols for detecting porcine reproductive and respiratory syndrome virus infection in boar studs by simulation modeling. J Vet Diagn Invest. 2007, 19:492-501.
- Salguero FJ, Frossard JP, Rebel JM, Stadejek T, Morgan SB, Graham SP, Steinbach F. Host-pathogen interactions during porcine reproductive and respiratory syndrome virus 1 infection of piglets. Virus Res. 2015, 202:135-43.
- Segalés J, Domingo M, Balasch M, Solano GI, Pijoan C. Ultrastructural study of porcine alveolar macrophages infected in vitro with porcine reproductive and respiratory syndrome (PRRS) virus, with and without Haemophilus parasuis. J Comp Pathol. 1998, 118:231-43.
- Sur JH, Cooper VL, Galeota JA, Hesse RA, Doster AR, Osorio FA. In vivo detection of porcine reproductive and respiratory syndrome virus RNA by in situ hybridization at different times postinfection. J Clin Microbiol. 1996, 34:2280-6.
- Terpstra C, Wensvoorst G, Pol JMA. Experimental reproduction of porcine epidemic abortion and respiratory syndrome (mystery swine disease) by infection with Lelystad virus: Koch’s postulates fulfilled. Vet Q. 1991, 13:131–36.
- Tian K, Yu X, Zhao T, Feng Y, Cao Z, Wang C, Hu Y, Chen X, Hu D, Tian X, Liu D, Zhang S, Deng X, Ding Y, Yang L, Zhang Y, Xiao H, Qiao M, Wang B, Hou L, Wang X, Yang X, Kang L, Sun M, Jin P, Wang S, Kitamura Y, Yan J, Gao GF. Emergence of fatal PRRSV variants: unparalleled outbreaks of atypical PRRS in China and molecular dissection of the unique hallmark. PLoS One. 200, 2:e526.
- Tong GZ, Zhou YJ, Hao XF, Tian ZJ, An TQ, Qiu HJ. Highly pathogenic porcine reproductive and respiratory syndrome, China. Emerg Infect Dis. 2007, 13:1434-6.
Van Alstine WG, Popielarczyk M, Albregts SR. Effect of formalin fixation on the immunohistochemical detection of PRRS virus antigen in experimentally and naturally infected pigs. J Vet Diagn Invest. 2002, 14:504-7. - Vézina SA, Loemba H, Fournier M, Dea S, Archambault D. Antibody production and blastogenic response in pigs experimentally infected with porcine reproductive and respiratory syndrome virus. Can J Vet Res. 1996, 60:94-9.
- Wensvoort G, Terpstra C, Pol JMA, Lask EA, Bloemraad M, de Kluyver EP, Kragten C, van Butten L, den Besten A, Wagenaar F, Broekhuijsen JM, Moonen PJM, Zetstra T, de Boer EA, Tibben AhJ, de Jong MF, van’r Veld P, Groenland GJR, van Gennep JA, Voets MTh, Verheijden JHM, Braamkamp J. Mystery swine disease in the Netherlands: the isolation of Lelystad virus. Vet Q. 1991, 13:121–30.
- Wills RW, Doster AR, Galeota JA, Sur JH, Osorio FA. Duration of infection and proportion of pigs persistently infected with porcine reproductive and respiratory syndrome virus. J Clin Microbiol. 2003, 41:58-62.
- Zimmerman JJ, Benfield DA, Dee SA, Murtaugh MP, Stadejek T, Stevenson GW, Torremorell M. Porcine reproductive and respiratory syndrome virus (porcine arterivirus). In: 10th ed. Diseases of swine, Ed. Wiley-Blackwell. 2012, 31:463-86.